-Search query
-Search result
Showing 1 - 50 of 583 items for (author: aly & n)

EMDB-28941: 
HIV Env BG505_MD39_B11 SOSIP boosting trimer in complex with B11_d77.7 mouse Fab and RM20A3 Fab
Method: single particle / : Torres JL, Ozorowski G, Ward AB

EMDB-28942: 
HIV Env germline targeting BG505_MD64_N332-GT5 SOSIP in complex with V3-glycan polyclonal Fab isolated from immunized BG18HCgl knock-in mice
Method: single particle / : Ozorowski G, Torres JL, Ward AB

EMDB-28945: 
HIV Env germline targeting BG505_MD64_N332-GT5 SOSIP in complex with V3-glycan polyclonal Fab isolated from immunized wild type mice, and NHP monoclonal Fab RM20A3
Method: single particle / : Ozorowski G, Ward AB

EMDB-43190: 
HIV Env BG505_MD39_B16 SOSIP boosting trimer in complex with B16_d77.5 mouse Fab and RM20A3 Fab
Method: single particle / : Ozorowski G, Torres JL, Ward AB
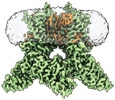
EMDB-42994: 
Apo-state cryo-EM structure of human TRPV3 in cNW30 nanodiscs
Method: single particle / : Nadezhdin KD, Neuberger A, Sobolevsky AI
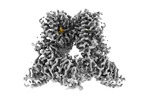
EMDB-42995: 
Open-state cryo-EM structure of human TRPV3 in presence of tetrahydrocannabivarin (THCV) in cNW30 nanodiscs
Method: single particle / : Nadezhdin KD, Neuberger A, Sobolevsky AI

EMDB-42996: 
Inactivated-state cryo-EM structure of human TRPV3 in presence of tetrahydrocannabivarin (THCV) in cNW30 nanodiscs
Method: single particle / : Nadezhdin KD, Neuberger A, Sobolevsky AI
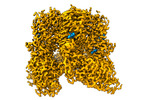

EMDB-42997: 
Open-state cryo-EM structure of human TRPV3 in presence of 2-APB in cNW30 nanodiscs
Method: single particle / : Nadezhdin KD, Neuberger A, Sobolevsky AI
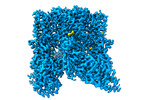

EMDB-42998: 
Inactivated-state cryo-EM structure of human TRPV3 in presence of 2-APB in cNW30 nanodiscs
Method: single particle / : Nadezhdin KD, Neuberger A, Sobolevsky AI

PDB-8v6k: 
Apo-state cryo-EM structure of human TRPV3 in cNW30 nanodiscs
Method: single particle / : Nadezhdin KD, Neuberger A, Sobolevsky AI

PDB-8v6l: 
Open-state cryo-EM structure of human TRPV3 in presence of tetrahydrocannabivarin (THCV) in cNW30 nanodiscs
Method: single particle / : Nadezhdin KD, Neuberger A, Sobolevsky AI

PDB-8v6m: 
Inactivated-state cryo-EM structure of human TRPV3 in presence of tetrahydrocannabivarin (THCV) in cNW30 nanodiscs
Method: single particle / : Nadezhdin KD, Neuberger A, Sobolevsky AI

PDB-8v6n: 
Open-state cryo-EM structure of human TRPV3 in presence of 2-APB in cNW30 nanodiscs
Method: single particle / : Nadezhdin KD, Neuberger A, Sobolevsky AI

PDB-8v6o: 
Inactivated-state cryo-EM structure of human TRPV3 in presence of 2-APB in cNW30 nanodiscs
Method: single particle / : Nadezhdin KD, Neuberger A, Sobolevsky AI

EMDB-43751: 
TRPM7 structure in complex with anticancer agent CCT128930 in closed state
Method: single particle / : Nadezhdin KD, Sobolevsky AI

PDB-8w2l: 
TRPM7 structure in complex with anticancer agent CCT128930 in closed state
Method: single particle / : Nadezhdin KD, Sobolevsky AI

EMDB-17375: 
Neisseria meningitidis Type IV pilus SB-GATDH variant
Method: helical / : Fernandez-Martinez D, Dumenil G

EMDB-17384: 
Neisseria meningitidis Type IV pilus SB-DATDH variant
Method: helical / : Fernandez-Martinez D, Dumenil G

EMDB-17386: 
Neisseria meningitidis Type IV pilus SA-GATDH variant
Method: helical / : Fernandez-Martinez D, Dumenil G

EMDB-17683: 
Neisseria meningitidis Type IV pilus SB-GATDH variant bound to the C24 nanobody
Method: helical / : Fernandez-Martinez D, Dumenil G

EMDB-17695: 
Neisseria meningitidis Type IV pilus SB-DATDH variant bound to the C24 nanobody
Method: helical / : Fernandez-Martinez D, Dumenil G

EMDB-17718: 
Neisseria meningitidis PilE, SB-GATDH variant, bound to the F10 nanobody
Method: helical / : Fernandez-Martinez D, Dumenil G

PDB-8p2v: 
Neisseria meningitidis Type IV pilus SB-GATDH variant
Method: helical / : Fernandez-Martinez D, Dumenil G

PDB-8p36: 
Neisseria meningitidis Type IV pilus SB-DATDH variant
Method: helical / : Fernandez-Martinez D, Dumenil G

PDB-8p3b: 
Neisseria meningitidis Type IV pilus SA-GATDH variant
Method: helical / : Fernandez-Martinez D, Dumenil G

PDB-8pij: 
Neisseria meningitidis Type IV pilus SB-GATDH variant bound to the C24 nanobody
Method: helical / : Fernandez-Martinez D, Dumenil G

PDB-8piz: 
Neisseria meningitidis Type IV pilus SB-DATDH variant bound to the C24 nanobody
Method: helical / : Fernandez-Martinez D, Dumenil G

PDB-8pjp: 
Neisseria meningitidis PilE, SB-GATDH variant, bound to the F10 nanobody
Method: helical / : Fernandez-Martinez D, Dumenil G

EMDB-40190: 
Local map of B3SB3L in complex with two-RBD-up state I of soluble SARS-CoV-2 Spike trimer
Method: single particle / : Liu WP, Shokr A, Mabrouk M, Aly N, Zhang J, Aschauer P, Gao HL, Selvaraj G, Elzoghby A, Chen B, Kawano T, Nasr ML

EMDB-19132: 
Structure of dynein-2 intermediate chain DYNC2I2 (WDR34) in complex with dynein-2 heavy chain DYNC2H1.
Method: single particle / : Mukhopadhyay AG, Toropova K, Daly L, Wells J, Vuolo L, Mladenov M, Seda M, Jenkins D, Stephens DJ, Roberts AJ

EMDB-19133: 
Structure of dynein-2 intermediate chain DYNC2I1 (WDR60) in complex with the dynein-2 heavy chain DYNC2H1.
Method: single particle / : Mukhopadhyay AG, Toropova K, Daly L, Wells J, Vuolo L, Seda M, Jenkins D, Stephens DJ, Roberts AJ

PDB-8rgg: 
Structure of dynein-2 intermediate chain DYNC2I2 (WDR34) in complex with dynein-2 heavy chain DYNC2H1.
Method: single particle / : Mukhopadhyay AG, Toropova K, Daly L, Wells J, Vuolo L, Mladenov M, Seda M, Jenkins D, Stephens DJ, Roberts AJ

PDB-8rgh: 
Structure of dynein-2 intermediate chain DYNC2I1 (WDR60) in complex with the dynein-2 heavy chain DYNC2H1.
Method: single particle / : Mukhopadhyay AG, Toropova K, Daly L, Wells J, Vuolo L, Seda M, Jenkins D, Stephens DJ, Roberts AJ

EMDB-41042: 
S. enterica WbaP in a styrene maleic acid liponanoparticle
Method: single particle / : Dodge GJ, Imperiali B

PDB-8t53: 
S. enterica WbaP in a styrene maleic acid liponanoparticle
Method: single particle / : Dodge GJ, Imperiali B

EMDB-41830: 
Lipidated recombinant apolipoprotein E4
Method: single particle / : Strickland MR, Rau M, Summers B, Basore K, Wulf II J, Jiang H, Chen Y, Ulrich JD, Randolph GJ, Zhang R, Fitzpatrick JAJ, Cashikar AG, Holtzman DM

EMDB-41831: 
Gradient-fixed lipidated recombinant apolipoprotein E4
Method: single particle / : Strickland MR, Rau M, Summers B, Basore K, Wulf II J, Jiang H, Chen Y, Ulrich JD, Randolph GJ, Zhang R, Fitzpatrick JAJ, Cashikar AG, Holtzman DM

EMDB-29423: 
Structure of Escherichia coli CedA in complex with transcription initiation complex
Method: single particle / : Liu M, Vassyliev N, Nudler E

PDB-8ftd: 
Structure of Escherichia coli CedA in complex with transcription initiation complex
Method: single particle / : Liu M, Vassyliev N, Nudler E

EMDB-15940: 
CryoEM structure of GroEL-ADP.BeF3-Rubisco.
Method: single particle / : Gardner S, Saibil HR

EMDB-15942: 
CryoEM structure of GroEL-GroES-ADP.AlF3-Rubisco.
Method: single particle / : Gardner S, Saibil HR

PDB-8ba8: 
CryoEM structure of GroEL-ADP.BeF3-Rubisco.
Method: single particle / : Gardner S, Saibil HR

PDB-8ba9: 
CryoEM structure of GroEL-GroES-ADP.AlF3-Rubisco.
Method: single particle / : Gardner S, Saibil HR

EMDB-40054: 
Cryo-EM structure of fish immunogloblin M-Fc
Method: single particle / : Lyu M, Stadtmueller BM, Malyutin AG

PDB-8ghz: 
Cryo-EM structure of fish immunogloblin M-Fc
Method: single particle / : Lyu M, Stadtmueller BM, Malyutin AG

EMDB-15939: 
CryoEM structure of nucleotide-free GroEL-Rubisco.
Method: single particle / : Gardner S, Saibil HR

EMDB-15941: 
CryoEM reconstruction of GroEL-ATP-Rubisco.
Method: single particle / : Gardner S, Saibil HR

EMDB-15943: 
CryoEM reconstruction of GroEL-GroES-ADP.AlF3-Rubisco, class I.
Method: single particle / : Gardner S, Saibil HR

EMDB-15945: 
CryoEM reconstruction of GroEL-GroES-ADP.AlF3-Rubisco, class III.
Method: single particle / : Gardner S, Saibil HR

EMDB-15946: 
CryoEM reconstruction of GroEL-GroES-ADP.AlF3-Rubisco, class IV.
Method: single particle / : Gardner S, Saibil HR
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
